Combining Machine Learning and Molecular Dynamics to Predict P-Glycoprotein Substrates
Por um escritor misterioso
Descrição

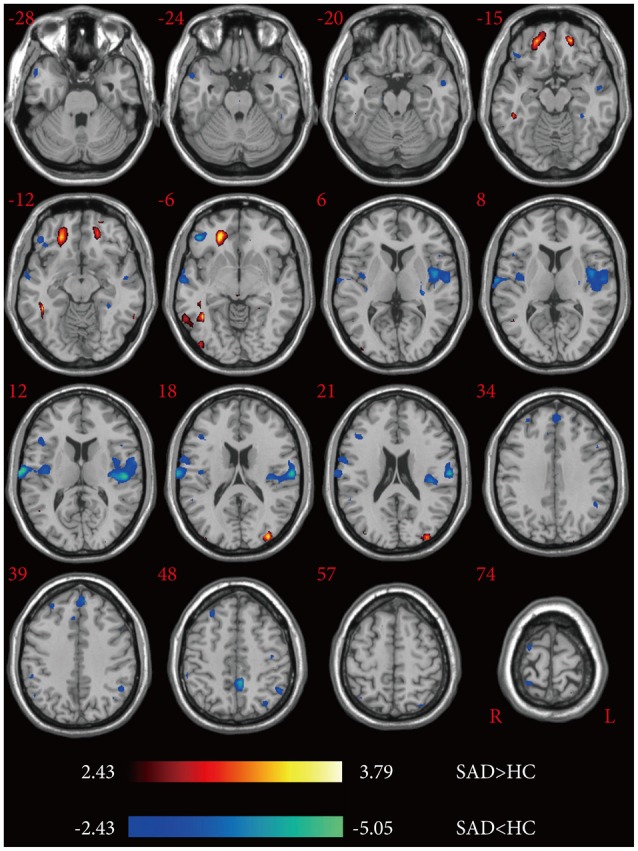
Structures of P-glycoprotein in different conformations (A) Domain

Interfacial Tension–Temperature–Pressure–Salinity Relationship for the Hydrogen–Brine System under Reservoir Conditions: Integration of Molecular Dynamics and Machine Learning
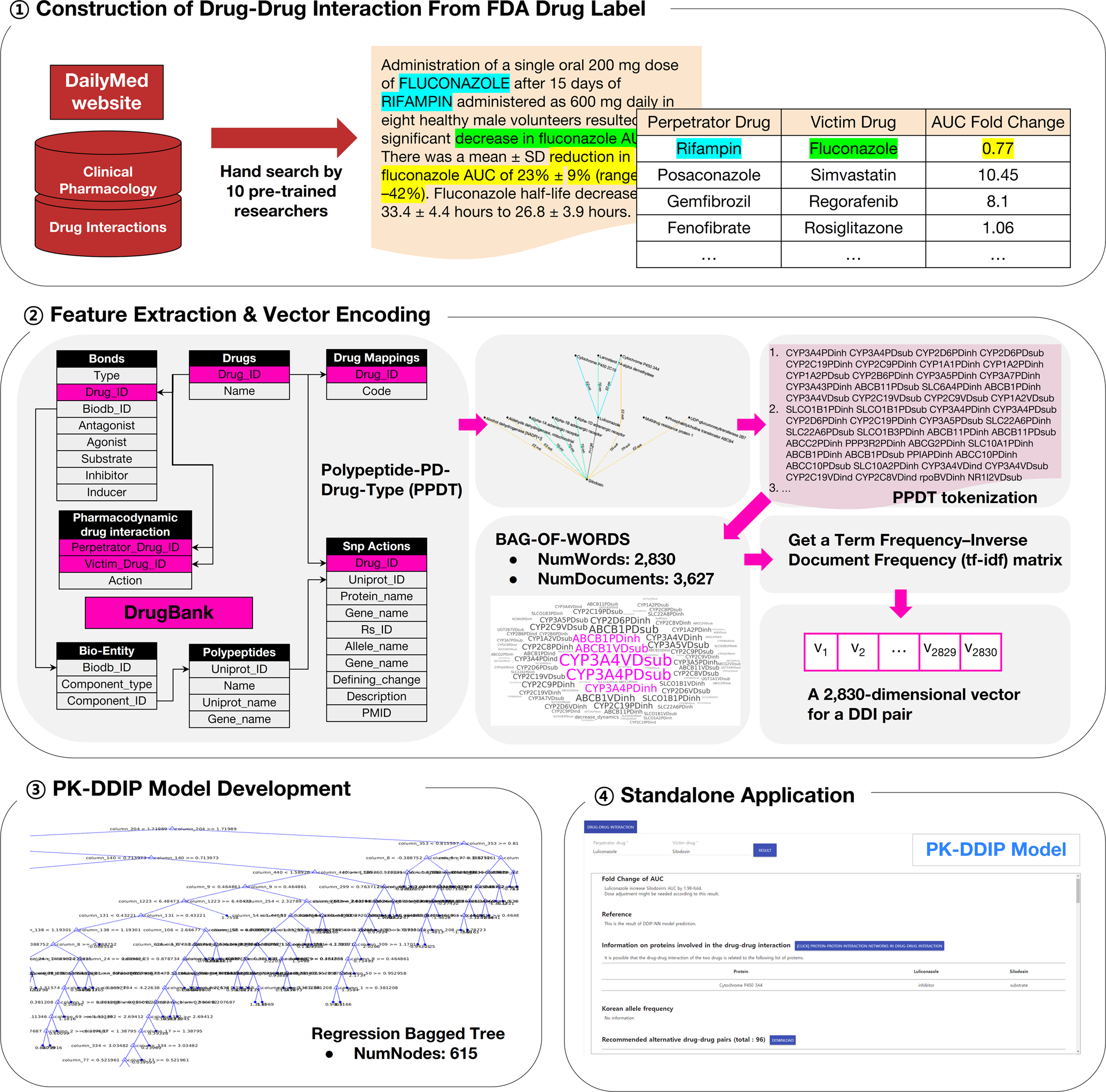

Machine learning-based quantitative prediction of drug exposure in drug-drug interactions using drug label information
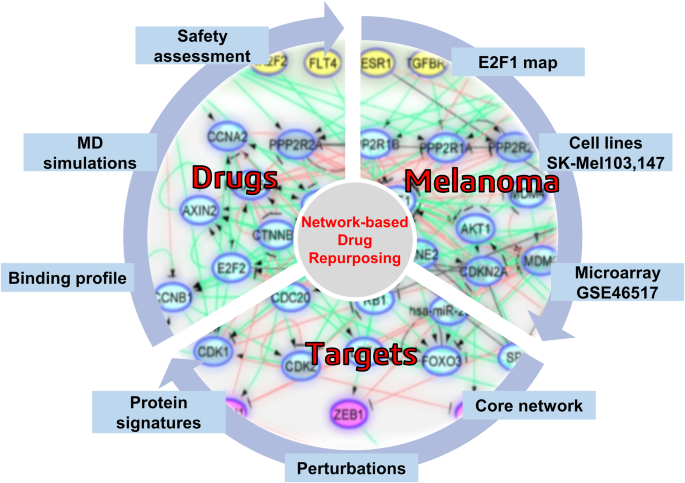
Logic-based modeling and drug repurposing for the prediction of novel therapeutic targets and combination regimens against E2F1-driven melanoma progression, BMC Chemistry
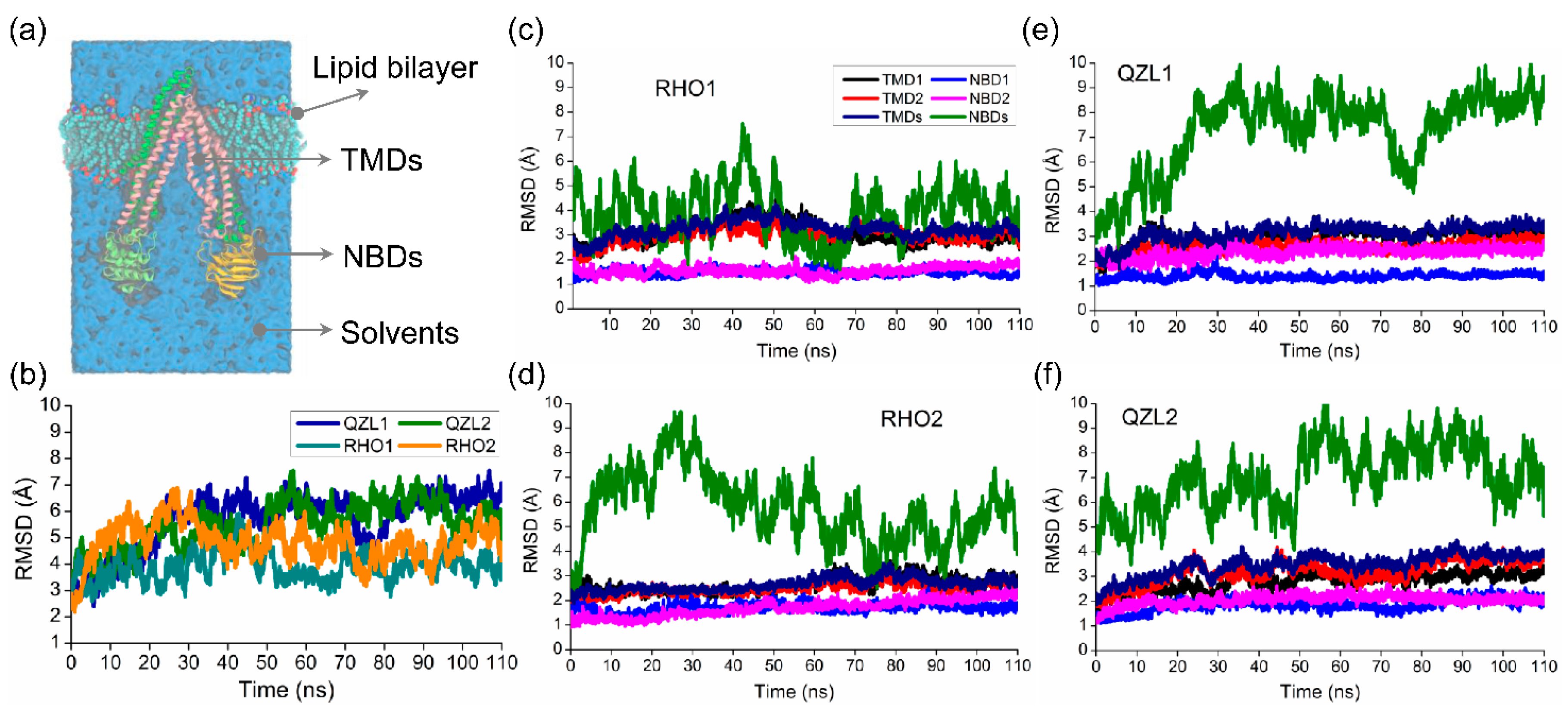
Molecules, Free Full-Text
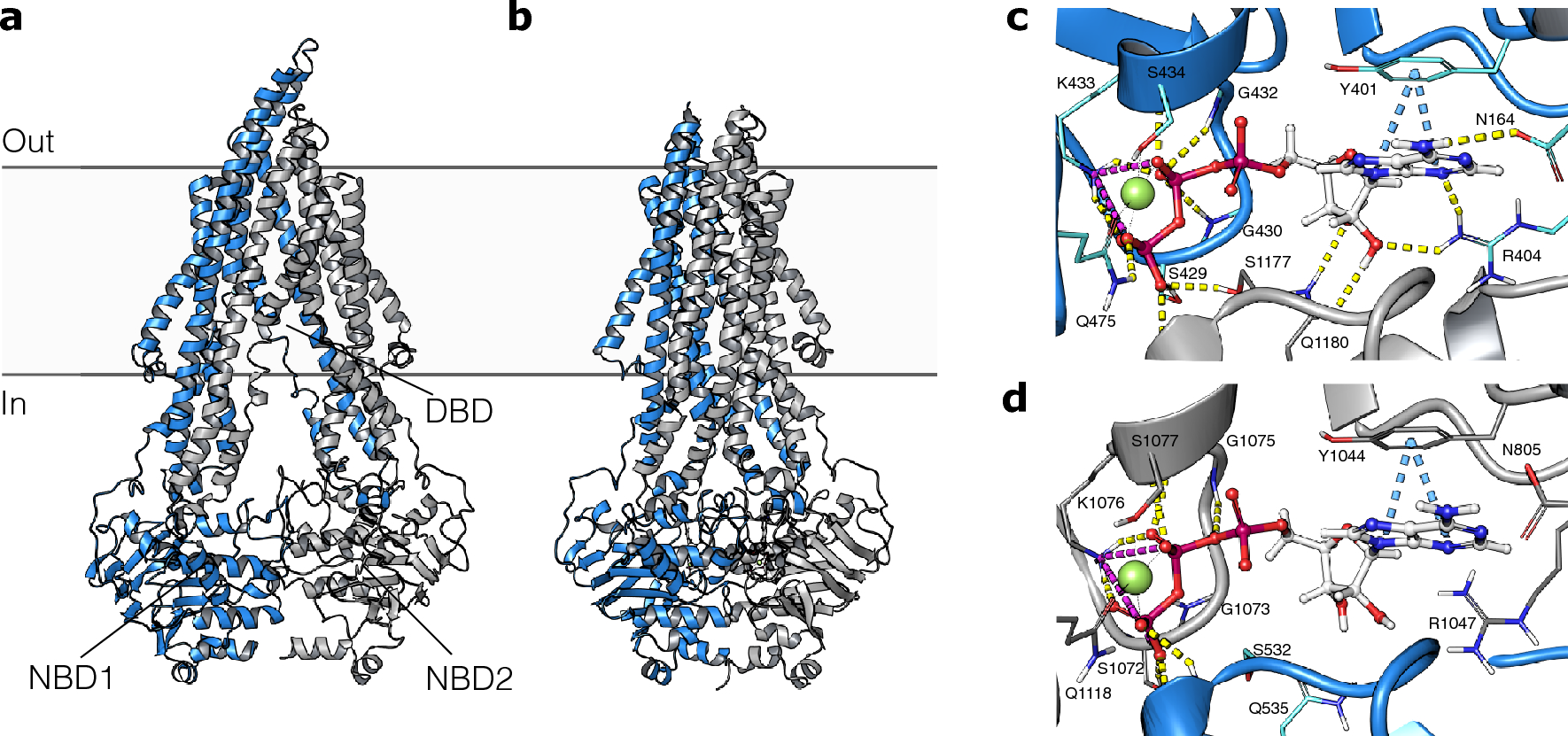
Structure-based discovery of novel P-glycoprotein inhibitors targeting the nucleotide binding domains

Novel QSAR Approach for a Regression Model of Clearance That Combines DeepSnap-Deep Learning and Conventional Machine Learning
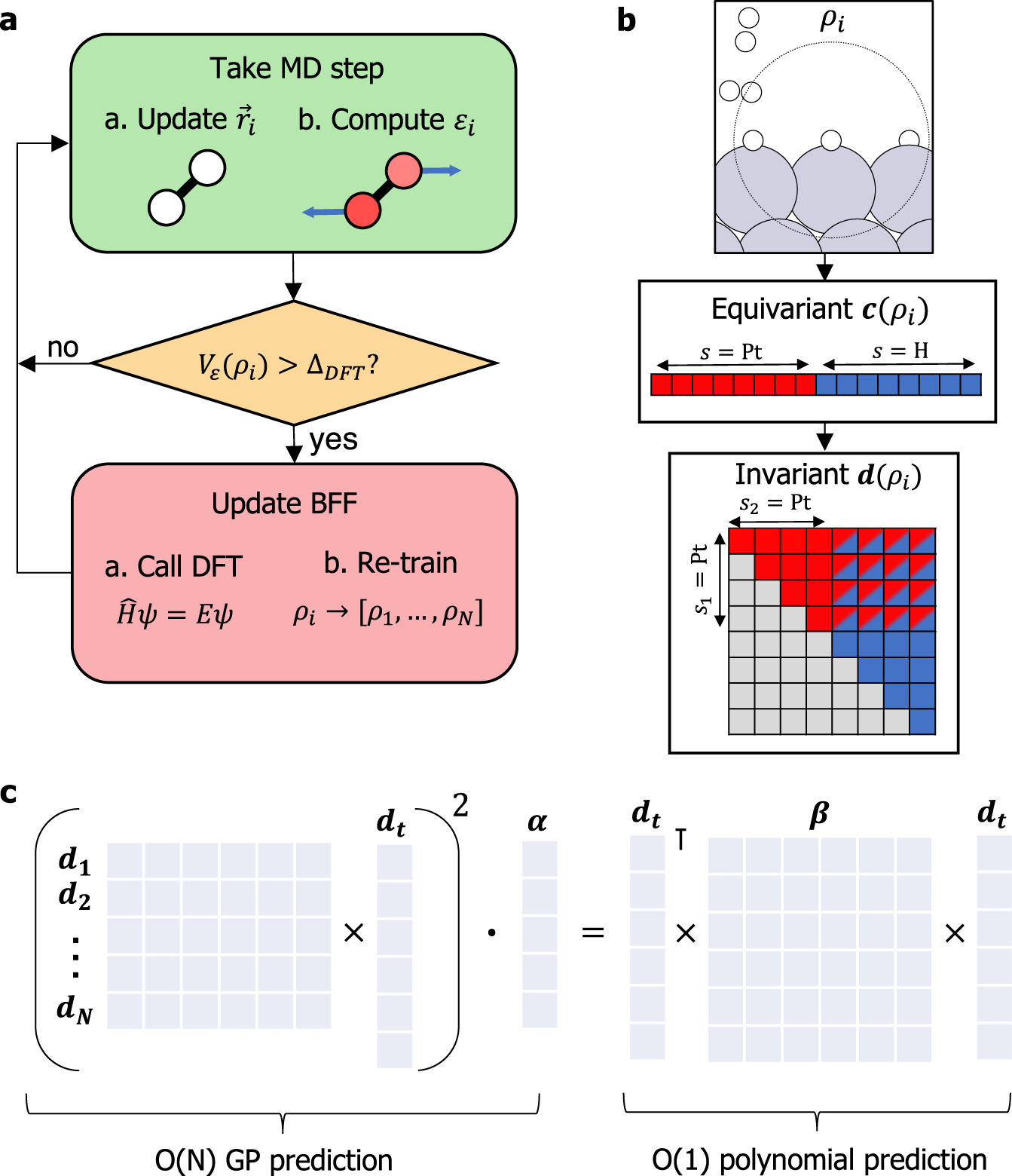
Active learning of reactive Bayesian force fields applied to heterogeneous catalysis dynamics of H/Pt

P-glycoprotein Substrate Models Using Support Vector Machines Based on a Comprehensive Data set

Computational and artificial intelligence-based approaches for drug metabolism and transport prediction: Trends in Pharmacological Sciences
de
por adulto (o preço varia de acordo com o tamanho do grupo)